Ethics statement
The YA, HA, PCI and AD samples were obtained from the University of Washington repository bank. In 2016, the University of Washington Institutional Review Board issued an official determination that the post-mortem repository work does not meet the metric of human subject research as this study collects samples from deceased individuals. These practices are now informed by the US Revised Uniform Anatomical Gift ACT 2006 (last revised or amended in 2009) and Washington Statute Chapter 68.64 RCW. Consent forms and HIPAA compliance are regulated by the University of Washington School of Medicine Compliance Office. All materials were collected under informed consent. For the SuperAgers, written informed consent and agreement to enter the study and the brain donation programme were obtained from all participants in the study, and the study was approved by the Northwestern University Institutional Review Board and in accordance with the Helsinki Declaration.
Donor cohorts and tissues
Where applicable, all human tissue studies were approved by the institutional review boards with oversight over the specific cohort studies, and all tissue samples were obtained with informed consent. Human brain tissue samples for molecular omics and sequencing studies were provided by the University of Washington (UW) BioRepository and Integrated Neuropathology (BRaIN) laboratory, which supports a number of cohort studies that were represented in the cohort for this study, including the UW Alzheimer’s Disease Research Center (ADRC) clinical core, the Kaiser Permanente Washington Health Research Institute Adult Changes in Thought (ACT) study, the Seattle Longitudinal Study (SLS) and the Pacific Northwest Brain Donor Network (PNBDN). Tissue samples were derived through a rapid tissue collection process, performed with a post-mortem interval of <12 h, and includes rapid procurement of the donor brain, coronal slicing and rapid sampling and freezing (either flash-freezing in liquid nitrogen or in supercooled dry ice and isopentane slurry), fixation of brain followed by routine sampling and diagnostic neuropathological analysis according to the National Institute on Aging and the Alzheimer’s Association guidelines for the pathological assessment of AD and related dementias44,45. This study was designed with five groups: YA, HA, PCI, AD and SA (Supplementary Table 1).
SA cohort
All participants were required to have preserved activities of daily living and to lack clinical evidence or history of neurological or psychiatric disease. The autopsied brains of participants characterized as cognitive SuperAgers from the Northwestern SuperAging Program were obtained from the Northwestern University Alzheimer’s Disease Research Center (NU-ADRC) Brain Bank. Written informed consent and agreement to enter the brain donation programme were obtained from all participants in the study, and the study was approved by the Northwestern University Institutional Review Board and in accordance with the Helsinki Declaration (https://www.wma.net/what-we-do/medical-ethics/declaration-of-helsinki/). Samples from representative brain regions of each participant were qualitatively surveyed and found to be free of significant neurodegenerative pathology other than common age-related changes. After autopsy, the right hemisphere was cut into 2–3 cm blocks and each block was flash-frozen on dry ice. Tissue at the mid-hippocampal region was collected from frozen blocks and used in this study. SA were required to be 80 years of age or older and to perform at or above average normative values for individuals in their 50s and 60s on the delayed recall of the Rey auditory verbal learning test (RAVLT)46, and within one standard deviation of the average range for their age and education on measures of other cognitive domains according to published normative values based on age, sex and race or ethnicity47,48.
Fresh-frozen preparation of samples and isolation of nuclei
The dentate gyrus was isolated from fresh-frozen blocks of 38 participants representing 5 diagnostic groups (YA, HA, PCI, AD and SA). Tissue was then immediately homogenized, whereby for each sample, a separate homogenizer and douncing pestles (loose and tight) were used. Each sample was homogenized in 1 ml pre-chilled lysis buffer (0.1% NP-40 alternative (or NP-40), 10 mM Tris, 146 mM NaCl, 1 mM CaCl2, 21 mM MgCl2 and 40 U ml–1 RNAse inhibitor) by mechanical douncing 20 times with the loose pestle followed by 20 times with the tight pestle to obtain a single-cell suspension. Cell suspensions were then incubated on ice for 5 min. The homogenate was then filtered through 70 µm filters (Miltenyi Biotec 130-041-407) and 40 µm filters (Miltenyi Biotec 130-041-406), moved to 1.5 ml tubes and centrifuged at 4 °C for 5 min at 500 rcf. Supernatants were removed and the nucleus pellet was washed 3 times with wash buffer (10 mM Tris, 146 mM NaCl, 1 mM CaCl2, 21 mM MgCl2, 0.01% BSA and 40 U ml–1 RNAse inhibitor). After the last wash, the supernatant was removed, and the nucleus pellet was resuspended in resuspension buffer and mixed with 900 µl Sucrose Cushion buffer. To remove additional debris, resuspended nuclei were loaded above a 3 Sucrose Cushion gradient (2.7 ml Nuclei PURE 2M Sucrose Cushion solution with 300 μl Nuclei PURE Sucrose Cushion buffer), and the sucrose gradient containing the nuclei was centrifuged at 13,000 rcf for 45 min at 4 °C. The supernatant was then carefully removed and the samples were immediately processed using the Single Cell protocol from 10x Genomics.
10x Genomics Multiome library preparation and sequencing
The 10x Genomics Multiome library preparation process and sequencing were done at Northwestern University NUseq facility core with the support of an NIH grant (1S10OD025120). The number of nuclei was analysed using Nexcelom Cellometer Auto2000 with the AOPI fluorescent staining method. The nuclei first underwent transposition with ATAC enzyme for 1 h at 37 °C. Next, 16,000 transposed nuclei were loaded into a Chromium Controller (10x Genomics, PN-120223) on a Chromium Next GEM Chip J (10x Genomics, PN-1000230), and single-cell gel beads were generated in the emulsion (GEM) according to the manufacturer’s protocol. Barcoded DNA and cDNA were PCR-amplified and subjected to library construction. The snATAC–seq library was generated using a Chromium Next GEM Single Cell Multiome ATAC + Gene expression kit (10x Genomics, PN-1000281) and Single Index Kit N Set A (10x Genomics, PN-1000212) according to the manufacturer’s instructions. Amplified cDNA was used for the gene expression library with a dual Index Kit TT Set A (10x Genomics, PN-1000215). Quality control for the constructed library was performed using an Agilent Bioanalyzer High Sensitivity DNA kit (Agilent Technologies, 5067-4626) and a Qubit DNA HS assay kit for qualitative and quantitative analysis, respectively. For the snATAC–seq library, the multiplexed libraries were pooled and sequenced on an Illumina Novaseq sequencer with 100 cycles kits using the following read length: 50 bp read 1 and 49 bp read 2. For the snRNA–seq library, the libraries were sequenced on an Illumina Novaseq sequencer with 100 cycles kits using the following read length: 28 bp read 1 for cell barcode and unique molecular identifiers (UMIs) and 90 bp read 2 for transcript expression. The targeted sequencing depth for snATAC–seq and snRNA-seq was 25,000 and 20,000 reads per cell, respectively.
10x single-cell multiome
Raw reads were demultiplexed and single-nucleus gene expression and peak enrichment were quantified simultaneously using CellRanger-arc count (10x Genomics). The quality of the run was assessed using the following criteria: (1) demultiplexing metrics, including the number of cells captured and the per cent of reads with valid barcodes; (2) gene expression metrics, including the per cent of mappable reads to the genome and transcriptome, and the median UMI counts and median genes expressed per cell; and (3) open chromatin metrics, including the per cent of mappable reads to the genome, in peaks and to promoter sequences, and the median counts and total peaks observed per cell. Following the quantification and peak calling analysis for each individual sample, all captures were aggregated to obtain a unified feature set for downstream analysis using CellRanger-arc aggr (10x Genomics).
Single-cell analysis
All samples were analysed together. Single cells were filtered to ensure that data used in the downstream analysis were of high quality. Cells with >10% mitochondrial expression, indicative of dead or dying cells, cells with low numbers of genes expressed (<1,000 genes) or low total UMI RNA counts (<2,000 UMI counts) and cells with low peak numbers (<200 peaks) or total ATAC counts (<500 counts) were removed. Clustering was performed on the RNA-seq data, anticipating that gene expression would have a higher dynamic range than open chromatin, using the Seurat package in R49. Single-droplet multiplets were detected using the union of two independent methods: Scrublet50 and DoubletDetection51. Gene expression was normalized using NormalizeData, and the top 6,000 variable genes were identified using FindVariableFeatures, both with default parameters. The top variable genes were z-scored using ScaleData, and principal components (PCs) were computed using RunPCA for the top 200 PCs. The significance of each PC was computed using JackStraw, and heatmaps of the top cells and genes per PC were plotted using DimHeatmap; after reviewing both results the top 125 PCs were selected as features for clustering analysis. Clustering analysis was performed with the Louvain algorithm as implemented in Seurat49 at resolutions 0.25, 0.5, 1 and 2. After reviewing the expression of known marker genes, we based our downstream analysis on the clustering results at resolution 1.
Cell-type determination
Primary cell types
To identify primary cell types, we used a transfer-learning approach based on scVI52 and scANVI28, in which reference dataset annotations are transferred to annotate subpopulations in a query dataset. Reference data were obtained from a previous study30, which reported a human developmental forebrain dataset, and another study of the human hippocampus11. The latent representation was modelled using the top 5,000 most variable genes in the combined dataset and was corrected for batch effect based on the data source. We then used scANVI (v.1.0.3)28 (n_samples_per_label=500) to transfer the most likely label from the reference datasets to the unannotated cells in our dataset. Based on the label transfer and gene marker annotations from clustering, we annotated astrocytes, neuroblasts, developmentally immature neurons, CA neurons, CA2–4 neurons, microglia, OPCs, mOLs, mGCs, endothelial cells and ependymal cells (Extended Data Fig. 6). NSCs were proposed to be part of the astrocyte cluster based on the recent literature53. Thus, astrocytes were provisionally labelled NSC/astrocyte.
Neuroblast and immature neurons
The primary cell-type analysis included cells identified as neuroblasts and developmentally immature neurons. To ensure the robustness of these annotations, we used previously published machine learning-based identification of immature neurons specifically in mGCs11. We then performed a subclustering analysis on cells identified as developmentally immature from the primary cell typing, which resulted in nine subclusters, and we performed a CytoTrace54 analysis to infer the relative maturity stage of each cell type. We observed a clear distinction between neuroblasts and immature neurons, as well as clear correspondence between subclustered cells with either neuroblast or immature neurons (Extended Data Fig. 7a,b). Subclusters 0, 2 and 6 were reclassified as neuroblasts and combined with the other neuroblast cells, and subclusters 1, 3, 4, 5, 7 and 8 were reclassified as immature neurons and combined with other immature neurons.
NSCs
To differentiate astrocytes from NSCs, we performed a subclustering of the NSC/astrocyte population and an RNA velocity analysis using scVelo55 on the NSC/astrocyte cluster along with neuroblasts, immature neurons and mGCs. Using the latent time measurement from the RNA velocity analysis, we identified a subcluster of the NSC/astrocyte population with the lowest latent time values (least differentiated) that was in between the remainder of the NSC/astrocyte cluster and the neurogenic cell types (Extended Data Fig. 1b,c). We identified these intermediate cells of NSC/astrocyte as the NSC subset, and the remainder as astrocytes.
Validation of the NSC signature
Systems analysis
Extended Data Fig. 1d,e and Supplementary Table 3 show the characterization of NSCs, neuroblasts and immature neurons based on the expression levels of developmental proxies (Extended Data Fig. 1d), the characterization of proxy expression trajectory in each cell type (Extended Data Fig. 1e) and open chromatin events of proxies in each cell type (Extended Data Fig. 1f). These were based on Supplementary Tables 5–7. We also computed differential gene and pathway analysis comparing NSCs to astrocytes. Pathway analysis revealed 57 pathways that were upregulated in NSCs, 23 of them are related to neuronal development, including axonal development, node of Ranvier, initial segment, growth cone, axonal guidance and dendritic spine (Supplementary Table 3).
We also compared NSC and astrocyte chromatin structure. We computed a UMAP of the ATAC–seq data of our NSC and astrocytes, which revealed that the NSCs have a chromatin structure that is distinct from astrocytes (Extended Data Figs. 1 and 4c).
Finally, SCENIC+ analysis was performed to study distinct GRNs (eRegulons) active in NSCs and mature astrocytes. This analysis also served as an orthogonal approach to assess whether the cells we annotated as NSCs represent a distinct population. This method integrates the chromatin accessibility and the expression of RNA at a single-cell level to infer the activity of TFs and to create GRNs. The regulon activity is an indicator of the activity of the TFs in GRNs. We found that NSCs and astrocytes formed distinct clusters based on the overall regulon embedding (Fig. 3a). This result further validates the annotation of NSCs as a distinct population. Diffusion analysis in the regulatory space of neurogenic cell types showed a clear developmental trajectory from NSCs to neuroblasts to immature neurons (Fig. 3b,c).
External comparisons
We compared NSCs in our dataset to one from a previous study13, which also performed single-cell sequencing of the human hippocampus. In the previous study13, the authors were able to identify both NSCs and neural progenitor cells. We identified the top 131 upregulated genes with adjusted P < 0.05 and log2[fold change] > 1 in the NSC/astrocyte population for our dataset and examined their expression levels in the other dataset13. Genes were selected only if they were expressed by >25% NSCs in both datasets. We observed that almost all of these genes were also upregulated in the NSCs of the previously published dataset relative to their astrocytes (Extended Data Fig. 2). This result illustrates a strong correspondence between the NSCs we identified and those independently obtained in the previous study13.
To enable putative identification of NSCs more broadly, we derived a NSC signature scoring metric based on the 131 NSC upregulated genes, computed as the mean z-score over all astrocyte or NSC cells. We computed NSC scores using previously published data13 and observed consistent high scores in their NSCs and low scores in their astrocytes (Extended Data Fig. 2). We also performed the reverse analysis, identifying upregulated genes in NSCs from the previously published data13 using the same criteria as described above. This analysis identified 161 genes, 55 of which were common to the 131 genes from our data. We then computed the mean z-score of these 161 genes both in our NSC/astrocyte group and in the cells from the previous study13 (Extended Data Fig. 2). We note that our NSCs also scored high using this approach, which again indicated a strong concordance in how NSCs are differentiated from astrocytes.
We next applied our annotation approach to a dataset that should not contain any significant number of NSCs. We chose a previously published sequencing dataset of the human prefrontal brain35. We would not expect any significant presence of NSCs in the prefrontal cortex. After performing label transfer, using our data as a reference and annotating cells from the prefrontal brain dataset35, a small number of apparent NSCs were labelled using this approach. However, when we examined the same 131 upregulated genes in NSCs in these cells, we observed a roughly even mix of upregulated and downregulated genes, which indicated that there is a lack of NSC signature and no defined population of NSCs (Extended Data Fig. 2).
Finally, we took a two-pronged approach and analysed two large datasets33,34 using two different methods. For the first dataset33, we examined each sample and tissue region, specifically those annotated as ‘astrocytes’, and used label transfer to predict NSCs. We reasoned that the similarity is highest between NSCs and astrocytes; therefore, populations that are typically labelled as astrocytes require in-depth analysis for possible subpopulations in the astrocyte cluster that might be NSCs. We did not observe any NSCs in any of the brain areas but did find a few modestly scoring cells in the pons for sample H19.30.002. Cells were labelled in red if their probability was >0.75 (Extended Data Fig. 7d). Label transfer is well-suited to broadly segregate and annotate cell populations based on a reference dataset. However, label transfer is not necessarily optimal for the identification of rare subpopulations, especially those that have high transcriptional similarity to larger subpopulations, as in the case for the large astrocyte population and the rarer NSC population. For the second dataset34, we used our targeted approach, looking specifically at overexpression of NSC-specific genes relative to astrocytes and considering a holistic annotation across the neurogenic spectrum. We performed a NSC scoring analysis using the above 131-gene signature in the cell population identified as astrocytes using ROSMAP. This analysis was performed for all six brain regions, independently for each sample to avoid sample-to-sample differences from skewing gene expression z-scores (Extended Data Fig. 8). No NSCs were observed in any of the other brain areas other than a few in the anterior thalamus. Further annotation of immature neurons and mGCs by label transfer (using our data as a reference, annotating cells in ROSMAP) in the samples with positive NSCs showed that we were able to identify a significant number of both immature neurons and mGCs in the hippocampus, but we did not identify immature neurons or mGCs in the anterior thalamus (Extended Data Fig. 7c). From the combination of these observations, we conclude that our use of the NSC scoring procedure is specific for identifying NSCs, and that further holistic analysis of neurogenic cell types—in particular the identification of the immature and mGC end points—is crucial for a final determination of the neurogenic niche.
Validation of the neuroblast signature
We performed an additional global validation of NSC and neuroblast signatures in a comparison a previously published dataset13. We computed DEGs of NSCs and neuroblasts, this time relative to all other cells in the tissue, for both our data and the previously published data13. We compared the number of common DEGs (log2[fold change] > 1, adjusted P < 0.05, per cent expressed > 25%) of similarly annotated cells (NSCs in both or neuroblasts in both) and computed the statistical enrichment using Fisher’s exact test. The overlap of DEGs for both NSCs and neuroblasts had large odds ratios and were highly significant. As a negative control, we considered the reverse comparison; that is, overlap of DEGs in NSCs in one dataset to those from neuroblasts in the other dataset; these were not significant and had odds ratios close to 1 (Supplementary Table 4). Together, these analyses show commonality in the gene sets in our NSC and neuroblast cell types compared with the previously published dataset13 and confirm that NSCs and neuroblasts are also distinct from each other.
We also performed additional specific comparisons to highlight differences between neuroblasts and oligodendrocytes, which have broadly similar transcriptional profiles. We show the expression levels of selected proxies of neuroblasts and oligodendrocytes in a dot plot (Extended Data Fig. 2a). We also computed DEGs and pathway enrichment of for neuroblasts versus mOLs. Pathway analysis revealed 169 pathways that were upregulated in neuroblasts. Of these, 80 pathways were directly related to axonal, dendritic and presynaptic and postsynaptic structure, function and plasticity (Supplementary Table 2). Finally, we performed an RNA velocity analysis over the population of neuroblasts and mOLs and observed a consistently lower latent time estimation for the neuroblast population, again reinforcing the relative level of differentiation of these cells (Extended Data Fig. 7a, b).
Peak recalling
After determination of cell types, we re-called peaks separately for each cell type using Seurat49, merging peaks across all cell types to obtain a more complete measurement of the open chromatin peaks for rarer cell types.
Cell-type abundance and statistical analysis
We counted the total cells per cell type per sample and computed association statistics between these cell counts and diagnosis of the participants and other AD-related traits using edgeR56 without the TMM normalization. Comparisons between groups, for example, cell abundance between AD and HA, were computed using the exactTest function56. Association of cell-type abundance with continuous variables, such as cognitive scores, were computed using generalized linear models. P values were adjusted for multiple testing using FDR correction.
Differential gene expression between clusters
DEGs for each neurogenic cell type were obtained using the FindAllMarkers function in Seurat49 with the Wilcox test, comparing each cell type to all other cells. This analysis was performed between NSCs, neuroblasts and immature neurons only. Significantly DEGs were determined based on adjusted q < 0.05.
Differential gene and peak expression by diagnosis
Differential gene and peak statistics between diagnosis groups were computed using a pseudo-bulk approach. Counts for gene expression or open chromatin were summed for each sample across all cells in a given cluster. Low-expressed genes or peaks—expressed in fewer than 25% of samples or with fewer than 50 total counts across all samples—were removed. Differential expression and open chromatin statistics for each cluster were computed using edgeR using the exactTest56 to perform pairwise between groups. P values were adjusted using FDR correction. DEGs were determined based on FDR < 0.05.
Motif analysis of differentially expressed peaks
Motif enrichment analysis enables inference of specific TFs that are driving developmental changes between clusters. First, we searched for instances of known TF motifs in all peak sequences from the JASPAR database57 using FIMO58. Then we computed motif enrichment statistics for each set of differentially expressed peaks by comparing the fraction of motif-containing peaks in or not in the differentially expressed peaks using Fisher’s exact test. We repeated this test for all motifs, correcting for multiple testing using the BH-corrected FDR59.
Pathway analysis of DEGs
Pathway enrichment of DEGs was interrogated against the Neuroimmune Gene Ontology (NIGO) Biological Process (BP) pathway database. Pathway enrichment for DEGs was performed with Fisher’s exact test in R. P values were adjusted for multiple testing using FDR correction.
GRN analysis
Regulatory interactions between TFs and target genes through open chromatin were determined using a similar TF–peak–gene trio approach to that as previously described39. First, low-expressed genes (expressed in <10% of cells) and peaks (observed in <2% of cells) were removed. Potential peak–gene interactions were identified by annotating each peak to all genes within 200 kb or overlapping the body of the gene. TF–peak interactions were identified by searching for TF motifs from the JASPAR core vertebrate database60 in open chromatin regions using FIMO61 with motif P < 1 × 10–5. Peak and gene log-normalized expression levels were averaged over the 20 nearest neighbours.
Potential regulatory interactions from TF gene–peak–target gene trios were constructed on the basis of TF–peak binding and peak–gene proximity in the genome, and Pearson’s correlation coefficients were computed on the normalized expression for each pair. TF gene–peak correlation coefficients less than zero were set to 0, and the inferred type of TF–target gene regulatory interaction (activation or repression) was based on the sign of the TF gene–target gene correlation. To rank and prioritize the interactions based on those with the strongest evidence, we computed the geometric mean of the absolute value of the three correlation coefficients. To scale this score with respect to the direction of regulation, we multiplied the score by the sign of the TF gene–target gene correlation.
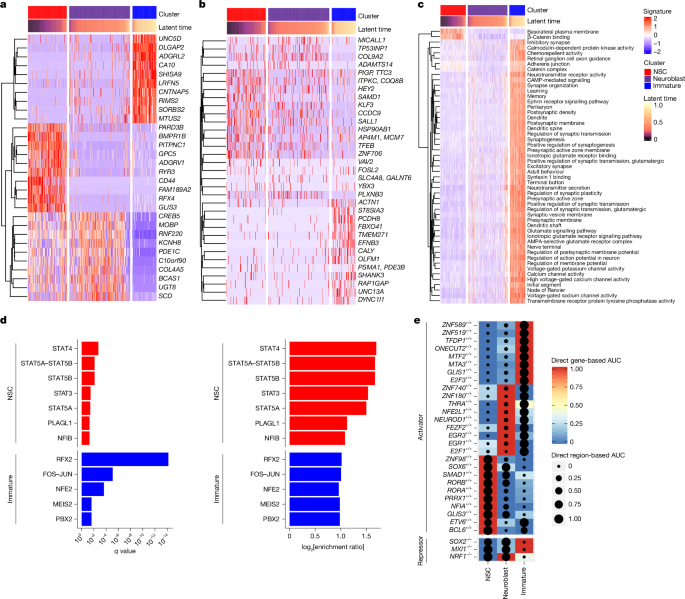
For GRN analysis of neurogenic cell types (NSCs, neuroblasts and immature neurons), we performed this analysis separately for cells from each diagnosis group (YA, HA, PCI, AD and SA) to enable comparisons of interaction scores between cell types. We also augmented the above-computed scores with the differential gene and peak statistics computed between cell types. For each cell type, we looked at the log fold change values in TF gene, peak and target gene expression. First, we verified that the sign of each correlation coefficient matched the sign of the product of the log2[fold change] values. Second, we computed the geometric mean of the absolute value of all three log2[fold change] values to obtain a cell-type-specific score.
eRegulon analysis to investigate GRNs
We ran SCENIC+ with region sets built from the 3,000 NTOP topics, OTSU topics, DARs between cell types and DARs between conditions in individual cell types. We used a custom cis-target database created with 1,000-base-pair padding from every region present in our filtered ATAC modality. Input motifs were taken from v10nr_clust-nr.hgnc. We generated these inputs separately for all cell types and neurogenic cell types. Both sets of inputs were independently processed through SCENIC+ using default parameters40.
Cell–cell interaction
Interactions between cell types were inferred using the CellChat62 package in R with default parameters for cells from astrocytes, CA neurons and neurogenic cells (NSCs, neuroblasts and immature neurons grouped together). An initial analysis was conducted over all cells from all diagnosis groups. A second analysis was conducted separately for each biological sample. In the second per-sample analysis, we quantified the same interactions that were identified in the first overall analysis to enable a fair comparison of interaction strengths from sample-to-sample. Differential statistics of the interaction probabilities was computed using limma in R63.
Resilience analysis
Specific filtering for ‘SA resilience’ genes and peaks was based on differential gene and peak expression by diagnosis results. AD–YA, AD–HA and SA–AD pairs were compared and then a summarized effect and significance statistic was computed as log2[fold change] × –log10[q]. All three comparisons were required to have the same sign (same direction). A summary score was computed as the geometric mean of the absolute value of all three summary statistics.
Healthy ageing analysis
Specific filtering for healthy versus unhealthy ageing, by comparing PCI and SA outcomes, was also based on differential gene and peak expression by diagnosis results. In this case, we looked for genes and peaks for which the SA–HA result is opposite to PCI–HA and PCI–YA, and that at least one of these comparisons had q < 0.2.
For CellChat healthy ageing interactions, we obtained interactions that were significantly altered in AD but displayed the same relative directionality in SA–HA, PCI–HA, and PCI–YA.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.